Damien Woods, Ho-Lin Chen, Scott Goodfriend, Nadine Dabby, Erik Winfree, Peng Yin
ITCS '13 doi: 10.1145/2422436.2422476 (2013).
Downloads: ITCS [PDF], 2 pages. Full article preprint from arXiv [PDF], 39 pages.
Abstract:
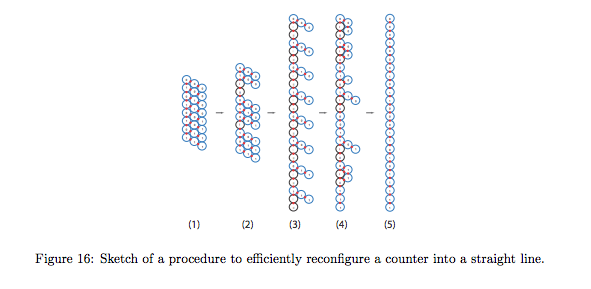
We describe a computational model for studying the complexity of self-assembled structures with active molecular components. Our model captures notions of growth and movement ubiquitous in biological systems. The model is inspired by biology's fantastic ability to assemble biomolecules that form systems with complicated structure and dynamics, from molecular motors that walk on rigid tracks and proteins that dynamically alter the structure of the cell during mitosis, to embryonic development where large scale complicated organisms efficiently grow from a single cell. Using this active self-assembly model, we show how to efficiently self-assemble shapes and patterns from simple monomers. For example we show how to grow a line of monomers in time and number of monomer states that is merely logarithmic in its length. Our main results show how to grow arbitrary connected two-dimensional geometric shapes and patterns in expected time polylogarithmic in the size of the shape plus roughly the time required to run a Turing machine deciding whether or not a given pixel is in the shape. We do this while keeping the number of monomer types logarithmic in shape size, plus monomers required by the Kolmogorov complexity of the shape or pattern. This work thus highlights the fundamental efficiency advantage of active self-assembly over passive self-assembly and motivates experimental effort to construct self-assembly systems with active molecular components.

Back to publications